Northshore Biosciences
Northshore Biosciences popped up on my radar again recently. There’s not a lot of information on the web, so I decided to skip forward in my list of DNA sequencing companies and write up a few notes on them.
Business
Northshore Biosciences was founded in 2009 as Lux Bio Group, Inc. [8] by Jonathan DeHart and Gordon Holt. A 2013 Genomeweb article states that an undisclosed series A was raised from Oregon Angel Fund and the ISB (Institute for Systems Biology?) [2].
A 2014 report by the Keiretsu Forum (a group of angel investors) states that they invested 21.2M USD in 35 companies during 2013 (including Northshore Bio). The average investment from Keiretsu was ~600K. SEC filings seem to indicate they’ve raise about 3M USD. Overall, it appears they’ve raise a few million and are at series A stage.
Technology
The Northshore Bio site doesn’t describe the technology in any depth, but it does show a nice video.
The fundamental Northshore approach is to create what they call “tuneable nanopores”. This is a fabrication approach where they create an aperture and then try and fill it in until they have a much smaller hole through which they can detect the translocation of bases. The genomeweb article suggests they are targeting 20nm long and 10nm wide nanopores.
It appears they are also proposing what they call “sequencing-by-degradation”. This is similar to the original Oxford Nanopore approach. Here an exonuclease, positioned near the aperture chews individual bases off a strand. These individual bases then go through the pore and are detected.
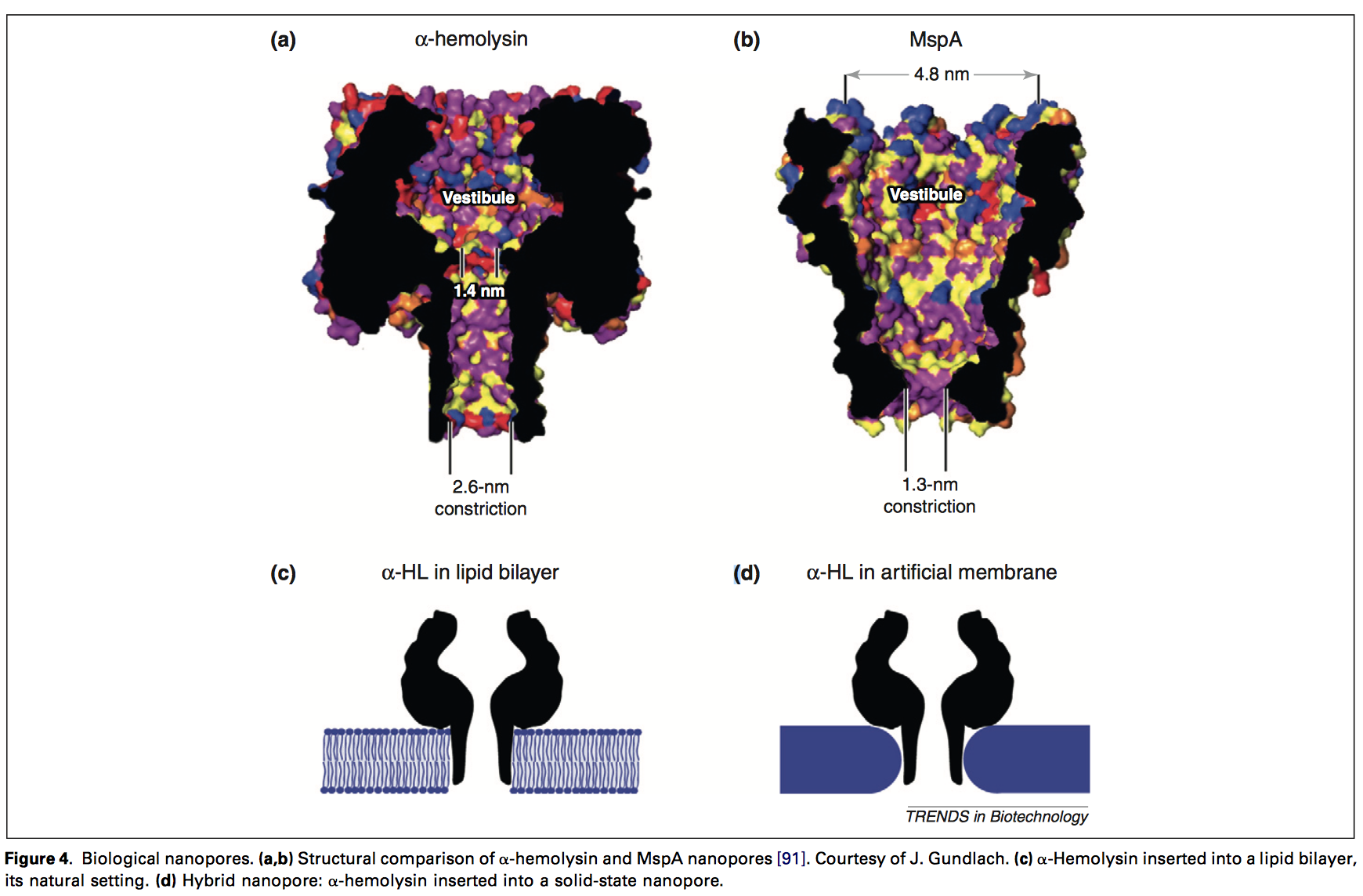
The pore size suggested (20nm by 10nm) is large when compared to typical protein nanopores, which typically have constrictions of 1 to 3nm:
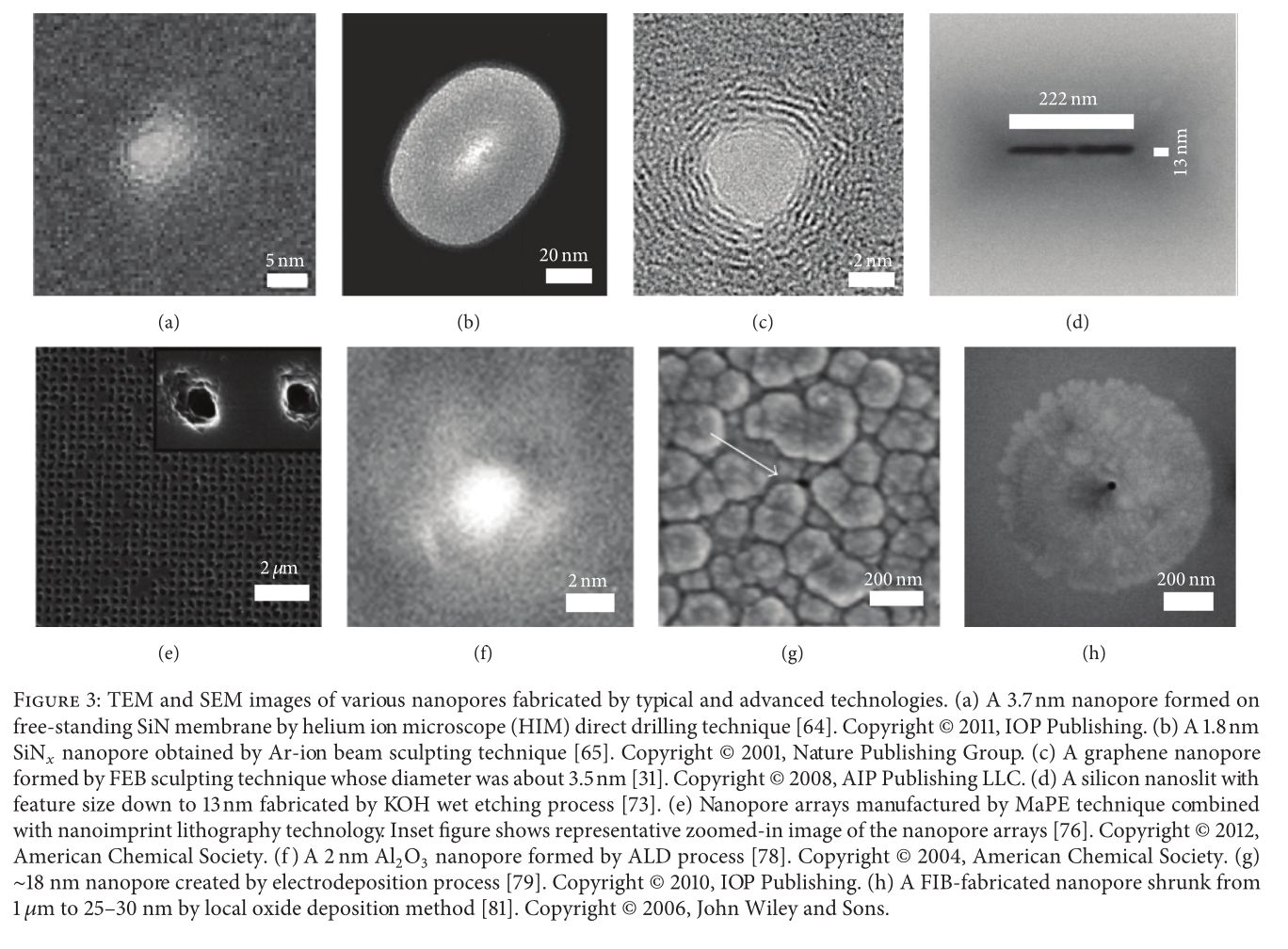
Solid state nanopores too, have achieved dimensions in the single digit nanometer range:
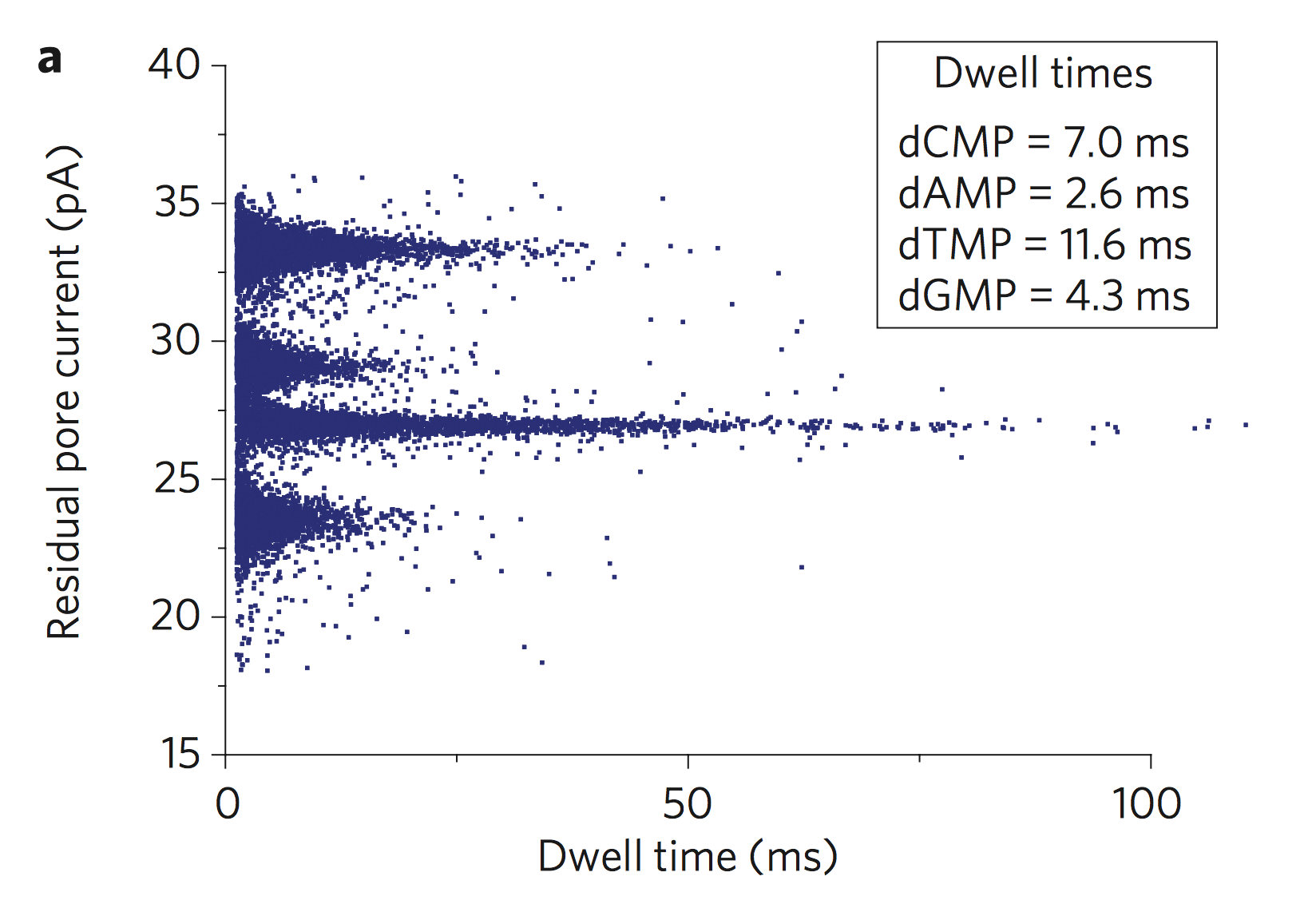
So it’s likely that the large dimensions of the pore somewhat motivate the decision to use an exonuclease, single nucleotide detection approach. One potential issue here is the dwell time of the bases in the nanopore. In many single base detection experiments the dwell time of bases is non-gaussian. There are many nucleotides that go through the pore so quickly that you can’t detect them:
In addition to the basic method shown on their site, Northshore Bio appear to have a number of patents (still assigned to Lux Bio Group Inc.) [1].
The patents discuss a number of different approaches. These include:
* Creating a small aperture, in which a bilayer just big enough for a single nanopore is formed (this could help with issues with multiple insertion of pores in other systems).
* Nanowells with sidewall electrodes.
* Nano-membranes which can change shape.
* Depositing membranes under electro-chemical control.
Most of the patents appeared to be continuations, and all authored by Gordon Holt. I looked through the patents for data (SEM images, or experimental data) and couldn’t find anything. It’s possible there’s more stuff in the pipeline.
The Northshore approach seems interesting, but not without significant challenges. I’d guess they will need significant funding to move forward (anything that involves nano fabrication does!). Will be watching with interest!
Notes
[1] Patent, Lux Bio Group: http://www.freepatentsonline.com/y2017/0298432.html
[2] Genomeweb article: https://www.genomeweb.com/sequencing/northshore-bio-develops-solid-state-tunable-nanopore-chips-sequencing-degradatio
“In late 2011, NorthShore Bio raised an undisclosed amount of funding in a Series A round with the Oregon Angel Fund and the ISB.”
“For sequencing applications, the company is targeting pore dimensions of less than 20 nanometers in length and 10 nanometers in diameter, similar to the nanopores explored for sequencing by others.”
“For sequencing, NSB is pursuing a sequencing-by-degradation approach, which is similar in principle to the exonuclease sequencing strategy Oxford Nanopore was exploring before it abandoned it in favor of DNA strand sequencing.”
[4] https://www.k4northwest.com/down/eJzLKCkpsNLXL87MyS4uSSwq0Ss21kvMTazKz0ssL9ZLzs%40VNzU2TjMyNDcB0pYG5gYphhYmZkaJpsZ6BSlpAJ08E30%3D/Keiretsu%20%20Forum%20Northwest%202013%20Funding%20Press%20Release.pdf
[5] https://doi.org/10.1016/j.tibtech.2011.07.006
[6] https://www.researchgate.net/publication/303696326_Solid-State_Nanopore-Based_DNA_Sequencing_Technology
[7] http://www.nature.com/articles/nnano.2009.12
[8] There’s still an old website online for Lux Bio Group: http://luxbiogroup.com/index.html
[9] https://www.whoisraisingmoney.com/lux-bio-group-inc