Lingvitae AS

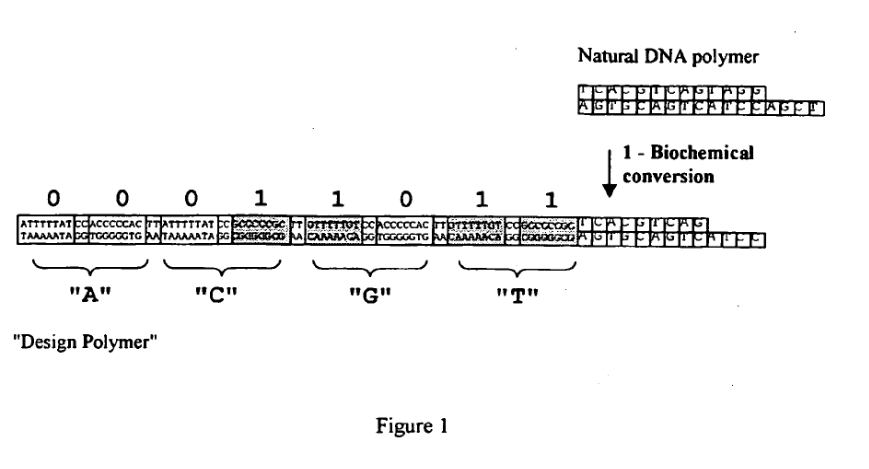
With the Stratos acqusition I wanted to write up a few notes on a much less well known (and now I think inactive) startup called LingVitae. LingVitae was a Norwegian single molecule sequencing startup. In many ways, their original “mission” was similar to Stratos’. They were working on a way to replace a single nucleotide with a magifying tag, or what is sometimes called a “Design DNA Polymer”.
The approach to generating these “expanomer-like” strands, is rather similar one approach suggested by Stratos in their patents. Essentially loop/hairpin like oligos are hybridized to the template and ligated:
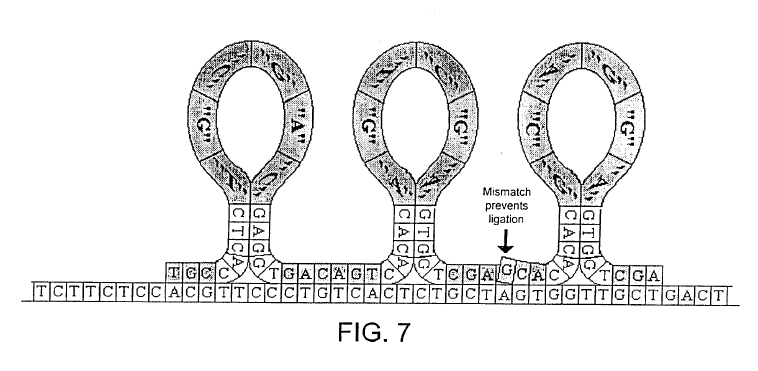
I’d guess there are a number of issues getting this to work. In particular, hybridization of such short (8mer) oligos might not be very specific. Will they really hybridize adjacently, specifically? Doesn’t the loop/bend cause a bunch of issues?
I couldn’t find anything that looked like experimental data in the patents. The closest I got was a 2007 paper describing the technique from Amit Miller [2]. This paper really just describes the concept and says “for proof-of-concept we designed and synthesized DNA oligonucleotides that encode”…”up to 8 bits of information”. Seems like they’re just synthesizing oligos, based on what LingVitae might have been able to produce. But there doesn’t seem to have been a followup paper showing any experimental work.
In their patents LingVitae, proposed a number of different read out methods, these include nanopore and optical approaches:
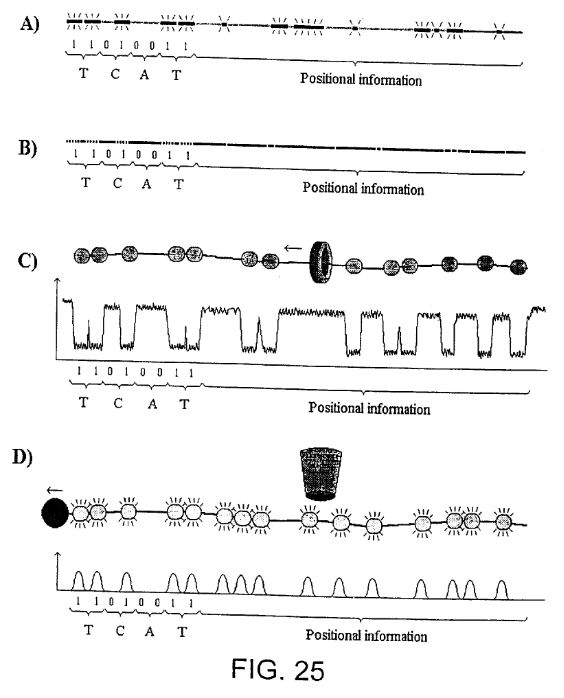
A) shows a DIRVISH based method using fluorescence labelled probes that bind the target molecules in a characteristic pattern,
B) shows an optical mapping based method in which the restriction pattern is used to give the position of the sequence,
C) shows a method in which a characteristic pattern of DNA binding proteins are registered as they pass through a micro/nano-pore and
D) shows a method using fluorescence labelled probes, proteins or the like which are registered as they pass a fluorescence detector.
LingVitae’s 2007 era website is shown below. This is from a time when they were patenting, and promoting work related to the design polymer idea. Their focus is on “high quality single molecule sequencing”.
This was before PacBio launched their instrument. So there were no “single molecule sequencing” platforms on the market at this time. LinkedIn shows about 30 former employees, so I imagine they had a reasonable team working on this.

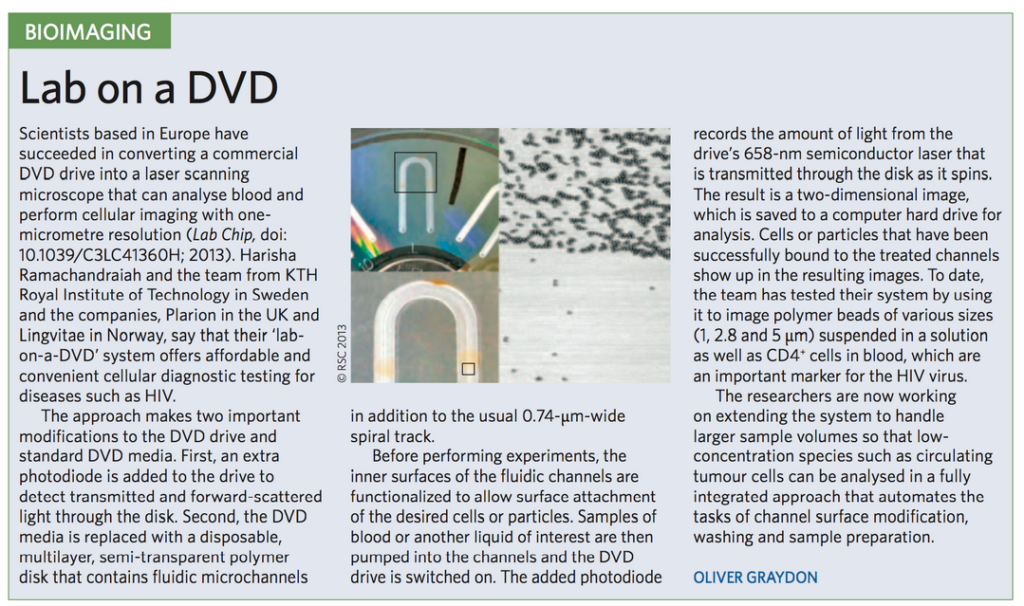
Slowly LingVitae seem to have moved away from DNA sequencing. The website, and PR shifted toward the development of a cheap cellular imaging platform [1]. The idea was to use a DVD drive as a microscope. This was always part of the sequencing play, but sequencing gets downplayed from 2012 onward:
www.lingvitae.com started redirecting to discipher.co sometime in 2017? But it seems like the site itself was offline before that… then sometime in 2014 the website went offline completely. In 2017 it started redirecting to discipher.co:

Which as of ~May 2019 also appears to be offline. LinkedIn doesn’t show any active employees so I assume this is the end of the LingVitae story.
It’s a shame really, it would have been interesting to see preliminary data from the design polymer approach. If anyone knows what happened to LingVitae please get in touch!
Notes/References
[1] https://www.theverge.com/2013/4/14/4223500/lab-on-a-dvd-blood-analysis-hiv-testing-fast-affordable
[2] Amit Miller paper: https://academic.oup.com/clinchem/article/53/11/1996/5627340
[3] Expandomer patent: https://patents.justia.com/patent/20090053699
Text from a version of their website, via the waybackmachine:
“Physical Magnification
The units to discriminate in a biological DNA molecule are bases or base pairs with a size of 0,34 nm each. The units to discriminate in a Design Polymer are blocks of up to 25 bases or base pairs with a size of up to 10 nm each.
Maximalisation of Unit Differences
The difference between the units to discriminate in a biological DNA molecule are only represented by a few atoms on a purine or pyrimidine skeleton attached to an identical backbone structure. The difference between the units to discriminate in a Design Polymer can be very significant and will be tailor made to achieve maximum resolution power on the read-out platform in question.
Binary Code
There are 4 units to discriminate between in a biological DNA molecule and the read-out platform must thus be able to distinguish between 4 different levels or states. There are 2 units to discriminate between in a Design Polymer and the read-out platform must thus be able to distinguish between only 2 different levels or states or alternatively even easier use a very simple on-off approach
Removal of secondary structures
A biological DNA molecule can take all forms of sequences and shapes and has a natural tendency to form secondary structures which can influence on the read-out process. A Design Polymer can be designed to avoid secondary structures and to ensure a reproducible behavior during the read-out process
Labels
It is difficult to label every individual base in biological DNA molecule due to sterical hindrance. The repertoire of labels that can be used is thus limited. Incorporation errors, quenching of neighbor labels, etc. adds to the challenge. It is easy to label every individual unit in Design Polymer as the spacing between labels, unit sequence, and more can be designed. The repertoire of labels that can be used are thus almost endless. Incorporation errors, quenching of neighbor labels, and other challenges can easily be solved by smart design of unit sequences, spacing, and other Design Polymer parameters.”
“The whole purpose of the Design Polymer Concept is to enable read out technologies to perform rapid DNA analysis with superior quality and resolution, and after years of extensive research LingVitae is now developing its first series of Design Polymer products -DNA EXPLORER SYSTEM.
DNA EXPLORER SYSTEM is going to be a lab-on-a-disk based kit designed to convert biological DNA into synthetic Design Polymers. The intention is to provide read out companies with a powerful tool that enables them to obtain advanced “DNA molecules” designed solely for the purpose of single molecule sequencing.
The first generation of the kit will consist of a conversion jig and a set of four conversion disks. It is intended to be very user-friendly, so that anyone with basic lab skills should be able to perform the conversion of nucleic acid in less than 24 hours. “
“DNA EXPLORER Fluidics Station 500
Fluidics Station 500 will be the 1st generation of fluidics stations for processing the Conversion Disks.
The Fluidics Station 500 will incorporate advanced design that provides improved ease-of-use and true walk away freedom to dramatically improve efficiency in the end-users genetic analysis. The system will run unattended until completion of a Conversion Disk, freeing the operator to attend to other responsibilities, thereby helping to improve the workflow and operation of the laboratory.
Total processing time per Conversion Disk will be 6 hours and the Fluidics Station 500 is designed to operate in environments running 2-4 daily runs per system. A total of 4 runs will be needed for whole genome conversion of a eukaryotic genome. No dedicated or special power requirements.”
“DNA EXPLORER Conversion Kit
The Conversion Kit will contain a series of 4 Conversion Disks (Disk 1-4) for the Conversion of 100 Gigabases of 24mers from a target material as well as reagents needed for DNA purification and initial handling of the sample material.
The Conversion Kit reagents will formulate into the fewest number of individual components possible, reducing preparation and handling steps. All of the reagents will be ready-to-use solutions.”
Video linked from site: https://www.youtube.com/watch?v=zLb9ip2lOos
Picture from their site, describing the “lab on a dvd”:
