Personal Genomics Inc./CrackerBio
 I recently came across a company called CrackerBio. They’d not come up on my radar before so I did some googling. As always, it’s worth noting that I have interests in this area and as such you should take my notes with a pinch of salt.
I recently came across a company called CrackerBio. They’d not come up on my radar before so I did some googling. As always, it’s worth noting that I have interests in this area and as such you should take my notes with a pinch of salt.
CrackerBio was formed in 2007 [1], their early press releases indicate they entered the Archon X Prize in 2009 (the prize turned out to be a bit of a flop really).
They’re an optical-semiconductor platform, as such it’s easy to make comparisons with Firefly. They are however quite different approaches. For a start, the CrackerBio system is single molecule.
The company seems to have undergone something of a reboot in the last year and is now called “Personal Genomics Inc.”. There’s a great video from a recent conference from which much of this post is sourced. There are also a number of patents, which I have only skimmed at present.
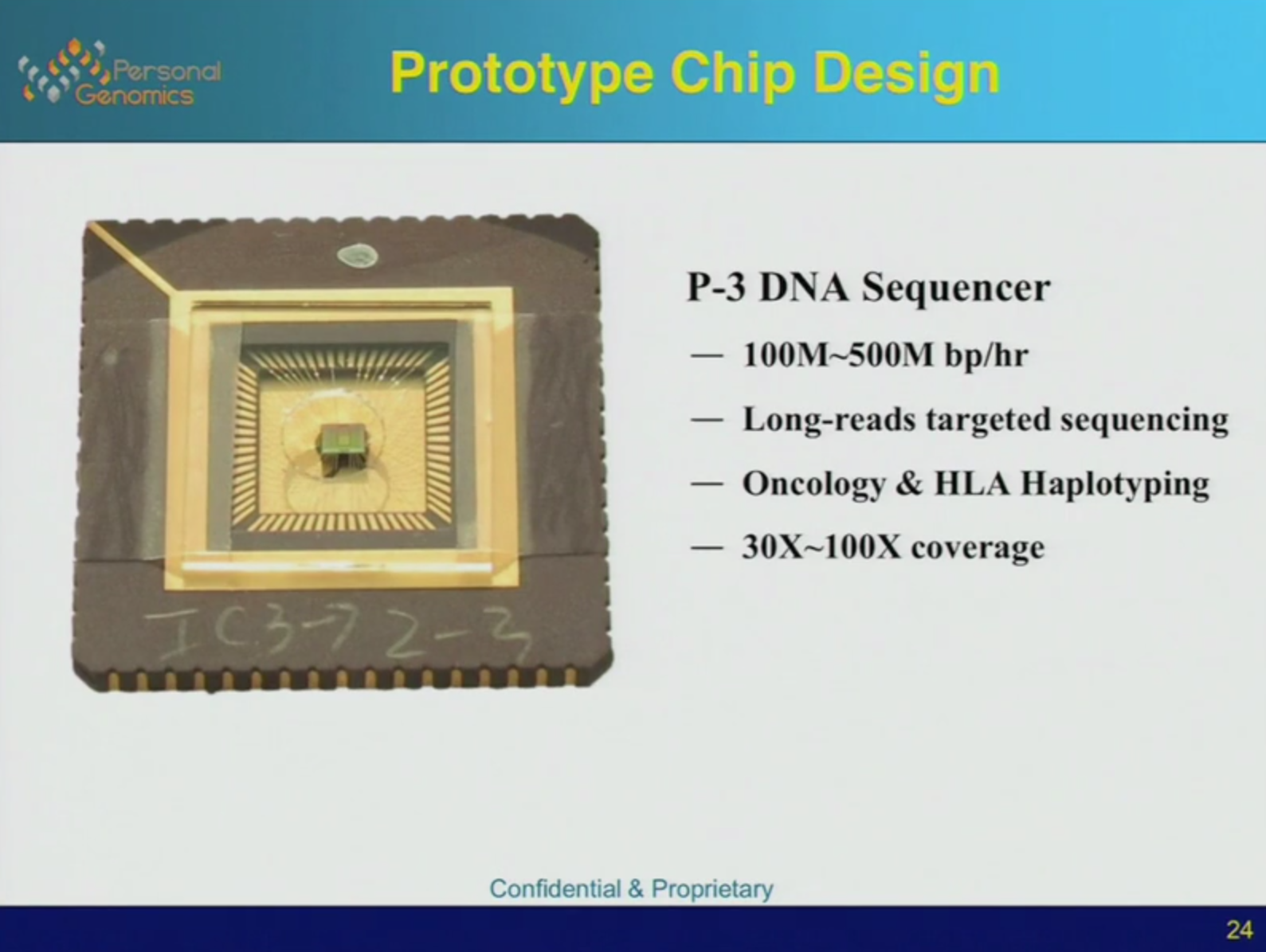
Anyway, here’s an overview. The prototype chip is shown below. They’re platform bares some similarity to PacBio’s. The incorporation of labelled bases on a strand sitting in a well is observed in real time. Unlike PacBio the whole system is integrated into a single chip.
The system looks quite neat, they plan to be to market in 2016. The CrackerBio site also showed 2015 as a launch date, and with semiconductor fabrication involved I’d guess there’s plenty of scope for delays.
Specs look quite reasonable, with estimates showing run throughput in the Gigabase range (much like Firefly). Unlike Firefly however they target long reads.
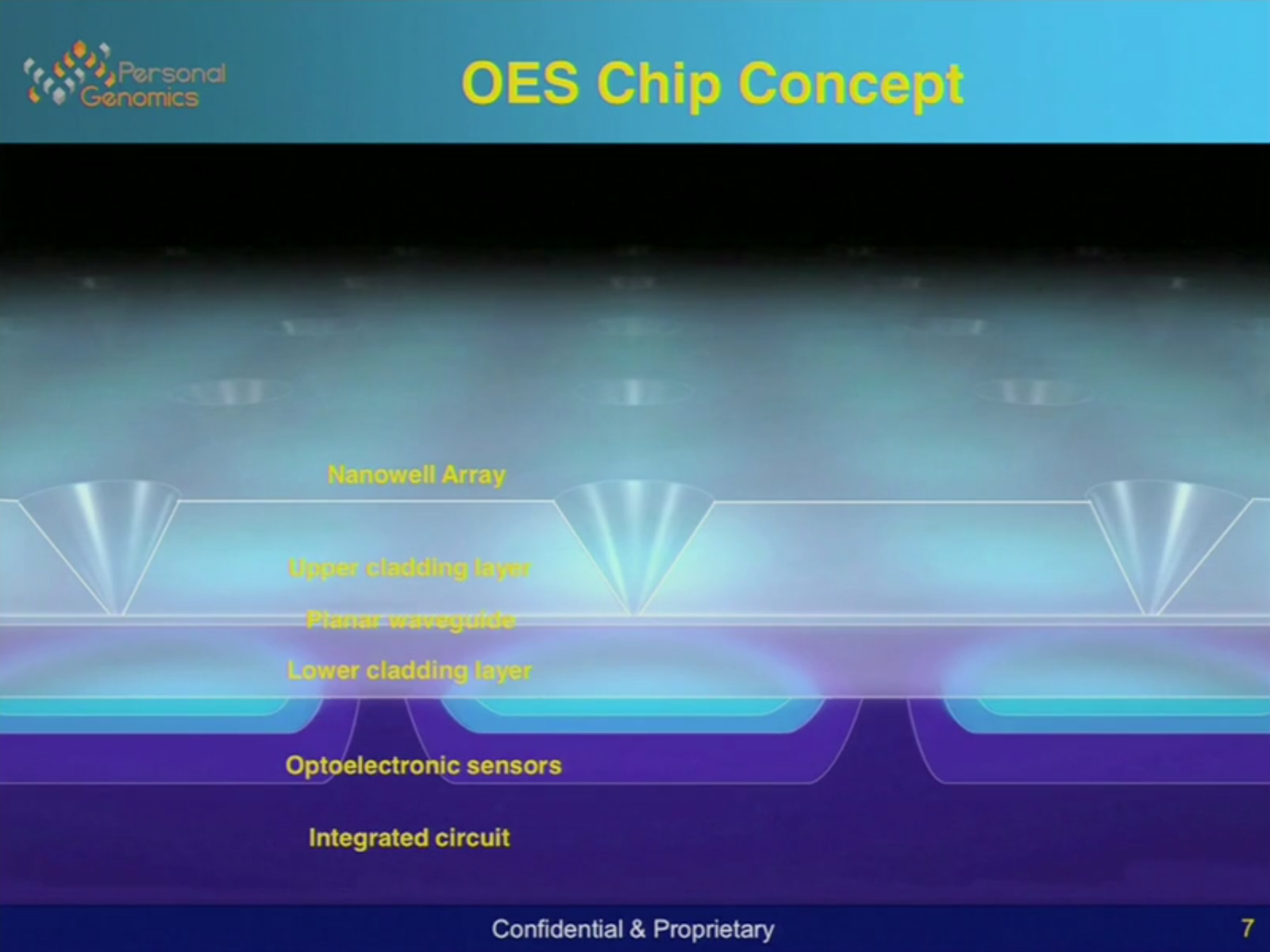
The system has a few neat aspects. A single molecule of DNA is attached to a bead, the beads sit in a nanowell. Under this well is a planar waveguide. My understanding is the waveguide is used to illuminate the bead. Planar waveguides in only one direction, so I guess this prevents the light from flooding into the sensor. The overall structure is shown below.
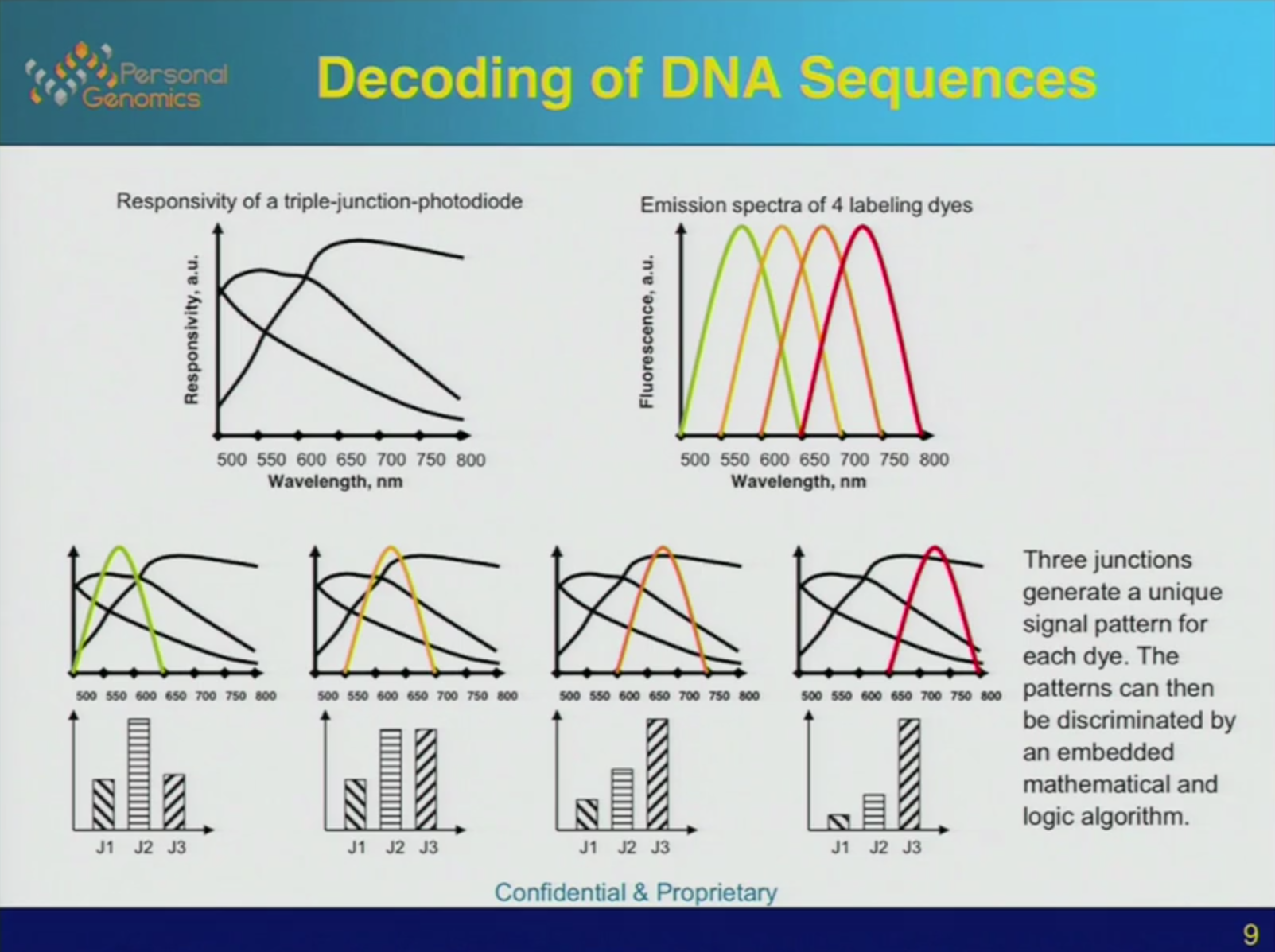
The final piece that makes the system viable is the triple-junction photodiode. Photodiodes are in general pretty sensitive to light across the spectrum. When we use a CCD camera or other photodiode array we usually combine them with filters. Either a fixed filter in scientific applications or a Bayer filter integrated into the device as in typical consumer cameras.
In the CrackerBio chip concept I don’t think it would really be viable to integrate filters into the device. So they take a different approach. Rather than using a single photodiode, they use a structure that effectively gives them 3 photodiodes, each with different response characteristics. Any one photodiode would not give a response current which would allow the frequency of light coming from the fluorescent label to be determined accurately. However the hope is that by combining the 3 different signals (shown to the left) an accurate determination can be made.
Overall I think the approach looks pretty neat, however realtime observation of the polymerase at work has proved problematic for PacBio and resulted in high error rates. From the looks of the system they will have one read per sensor, this would limit throughput. Having to spin a chip to advance the platform, also brings its own issue (takes money and time). It’s an interesting concept however, and I plan to keep an eye on their progress.
[1] From here. ABOUT TEAM CRACKER -Initiated in 2007 with support from the Industrial Technology Research Institute (ITRI), team cracker is based in Hsinchu, the heart of the Taiwanese “Silicon Valley.” Under the direction of Dr. Chung-Fan Chiou, an engineer, innovator and business developer, team cracker blends the expertise of young, highly talented and innovative opto-electronics engineers, organic chemists and molecular biologists. The team is developing a new ultra-high speed, low cost and portable method to sequence long base-pair reads with high accuracy. It is based on the sequential conversion of photons to electrons and to DNA sequence, on a single composite chip. The versatility of the underlying design is expected to revolutionize studies of biomolecules at the single-molecule level. For more information, please visit www.crackerbio.com.