INHECO 96 Chick Brooder
Overview
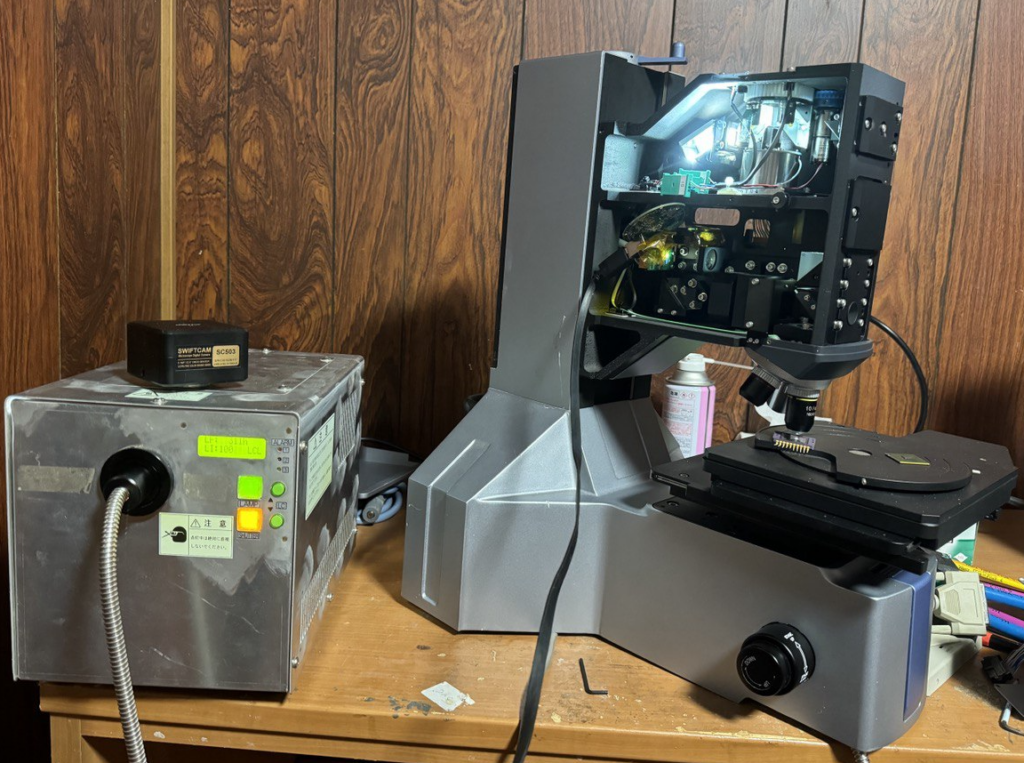
The Inheco Control 96 is an OEM unit built around a Watlow Series 96 PID controller and a Watlow LSTW driver module, housed in an Inheco-branded chassis. Originally designed for TEC/Peltier temperature control in laboratory equipment (e.g. the Inheco CPAC Ultraflat for heating Eppendorf tubes), it can be repurposed as a general-purpose temperature controller.
This writeup documents repurposing one such unit to drive a Kotatsu heater element (~100V rated, operated at 30–40V) as a brooder heater for day-old chicks. The unit was acquired second-hand and arrived with a PIC microcontroller attached to the RS232 port and the controller in a misconfigured/locked state.
Reference also: 41j.com Inheco Control 96 Notes — confirms the LSTW is an undocumented OEM module, likely an H-bridge in the original TEC application, but in single-output configurations acts as a simple DC power switch.
Hardware
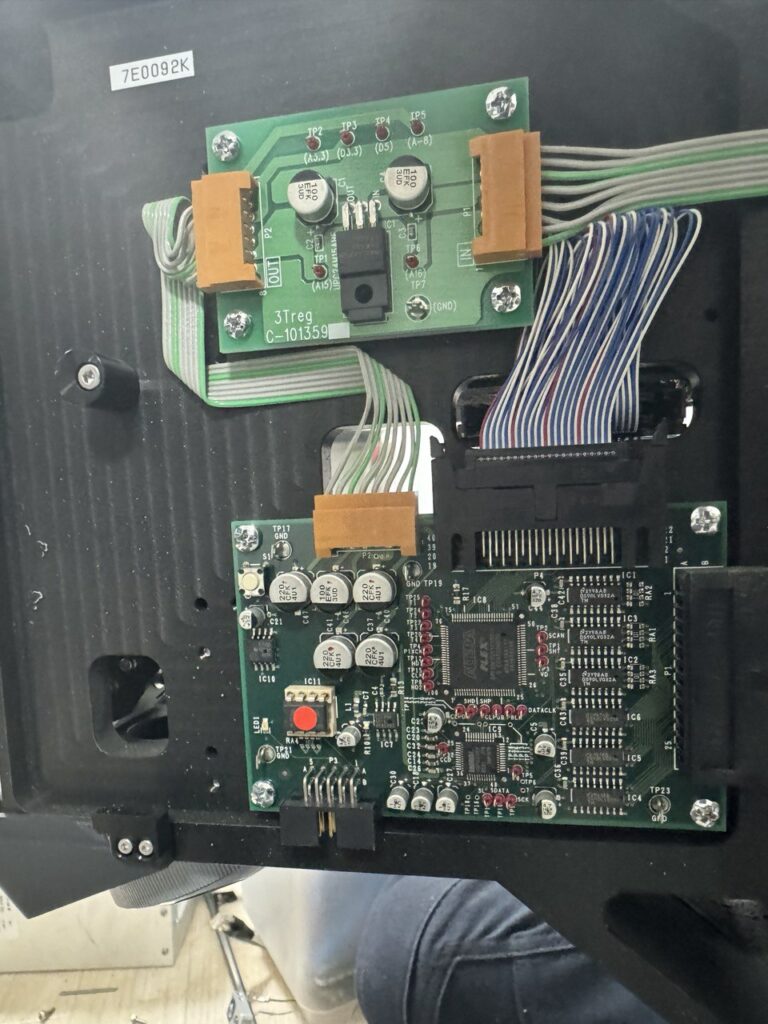
Watlow Series 96 PID Controller
- 1/16 DIN panel mount PID controller
- Universal input (thermocouple, RTD, process)
- 4 outputs (configuration dependent on model)
- RS232 serial communications on output 4 terminals (19, 20, 21)
- Keys: ▲ Up, ▼ Down, ∞ Home/Infinity, ↺ Advance/Cycle
- Firmware version on this unit: r7dL (visible in DIAG menu)
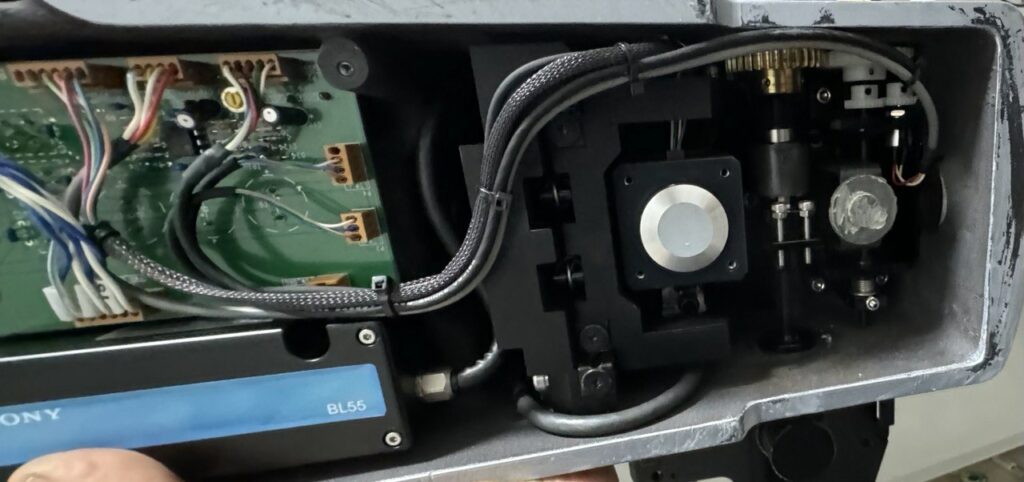
Watlow LSTW 1.5 Driver Module
- Undocumented OEM module, not available on Watlow website
- 8-terminal connector block
- Functions as a DC power switch (not a true SSR)
- Contains an IRF5305 P-channel MOSFET (55V, 31A rated) as the main switching element
- Protection diode across input: originally B13 Schottky (30V, 1A — undersized)
- Status LEDs: GRUN (green) = control input active, ROT (red) = output switching on
LSTW Terminal Pinout (as determined empirically)
| Pins | Function |
|---|---|
| 1–2 | Unknown / unused in this config |
| 3–4 | Output to load (heater) |
| 5–6 | Control input from Watlow 96 (24V DC signal) |
| 7–8 | Connected to Watlow 96 rear center terminals (power/common) |
The LSTW passes the supply voltage through to the load when the control signal is active — it does not have a separate mains input. Supply voltage = output voltage.
Kotatsu Heater Element
- Rated 100V AC
- Operated at 30–40V DC (sufficient for brooding temperatures)
- Connected to LSTW output terminals 3–4
Thermocouple
- Type J (blue/red wire, European IEC colour code)
- Connected to Watlow 96 Input 1 terminals
- Polarity sensitive — blue wire to negative terminal
Watlow Series 96 — Key Navigation
| Action | Key Sequence |
|---|---|
| Home Page | Press ∞ briefly |
| Operations Page | Press ▲ + ▼ together (~3 sec) |
| Factory Page | Hold ∞ + ↺ together for 6 seconds |
| Setup Page | Hold ▲ + ▼ together for 6 seconds |
| Enter a menu | Press ↺ |
| Change value | ▲ / ▼ |
| Confirm value | Press ↺ |
Some config settings appeared to be locked out at first. I’m not sure if I was just misunderstanding the config system or if I actually fixed this. There was a PIC connected to the RS232 input on the Watlow 96, this seemed to be processing a signal from a knob on the front of the unit. I disconnected this.
Configuring the Thermocouple Type
Accessing CIN1 (Calibration Input 1 Menu)
- Enter Config Page: hold ▲+▼ for 6 seconds (it changes to one menu after 3s then again)
- Press ▼ to navigate to CIN1
- Press ↺ to enter
- Set the thermo couple type to H to K-type and J for J-type.
LSTW Diode Failure and Repair
What Happened
When attempting to supply ~40V DC to the LSTW input (to drive the Kotatsu heater at sufficient power), the protection diode B13 failed with visible smoke.
Root Cause
The B13 is a 30V, 1A Schottky diode — it is connected across the input terminals as a flyback/protection diode. Supplying 40V exceeded its 30V rating, causing it to fail.
The main switching MOSFET (IRF5305, P-channel, 55V/31A, TO-220 package) survived.
Repair
The blown B13 was removed. Unit was tested without it — functional short-term since the MOSFET is not switching at high frequency in this application, so flyback spikes are minimal.
Replacement diode found by scavenging: A D2S58 Schottky diode was recovered from scrap PCB.
D2S58 specifications: 80V, 2A — significantly better rating than the original B13 and well suited for this application.
Installation: Cathode (stripe) to positive input terminal. Solder across input terminals in same orientation as original B13.
The D2S58 replacement is an improvement over the original B13. If sourcing a new part, any Schottky diode rated ≥50V, ≥1A is suitable (e.g. 1N5819 at 40V is marginal; prefer SS34, 1N5822, or similar at 40V+).
Final Working Configuration
| Parameter | Value |
|---|---|
| Thermocouple type | J type (J in CIN1) |
| Thermocouple wiring | Blue = negative, Red = positive |
| LSTW control signal | 24V DC from Watlow 96 Output 1 (terminals top 3-pin block) |
| Supply voltage to LSTW | ~40V DC (Rigol DP832) |
| Heater | Kotatsu element, connected to LSTW output terminals 3–4 |
| LSTW protection diode | D2S58 (scrap), across input, cathode to positive |
| Setpoint | Set to desired brooding temperature (e.g. 35°C for day-old chicks) |
Operational Notes
- The GRUN LED on the LSTW indicates the 24V control signal is present and active
- The ROT LED indicates the output is switching (heater powered)
- If ROT is lit but no heat: check supply voltage is connected to LSTW and heater is connected to output terminals 3–4
- ERR4 on the Watlow 96 = open sensor (thermocouple disconnected or broken) — check wiring at terminals 5, 6, 7
- The controller PID will modulate output to maintain setpoint — allow time for auto-tuning to stabilise temperature control
- A few degrees offset between reading and actual is normal with replacement thermocouples — a calibration offset can be added in the IN1 menu if accessible